Module 2: R and Tidyverse Foundations
Conducting Exposome-Wide Association Studies
Overview
In this module, we will cover:
- The pipe operator (
%>%and|>) - Six core dplyr verbs:
filter,select,mutate,arrange,summarize,group_by - Recoding variables with
case_when - Tidy data and reshaping with
pivot_longer/pivot_wider - Visualization with ggplot2
- Linear regression in R and the broom package
- Running many regressions with
nest_by
The Tidyverse Ecosystem
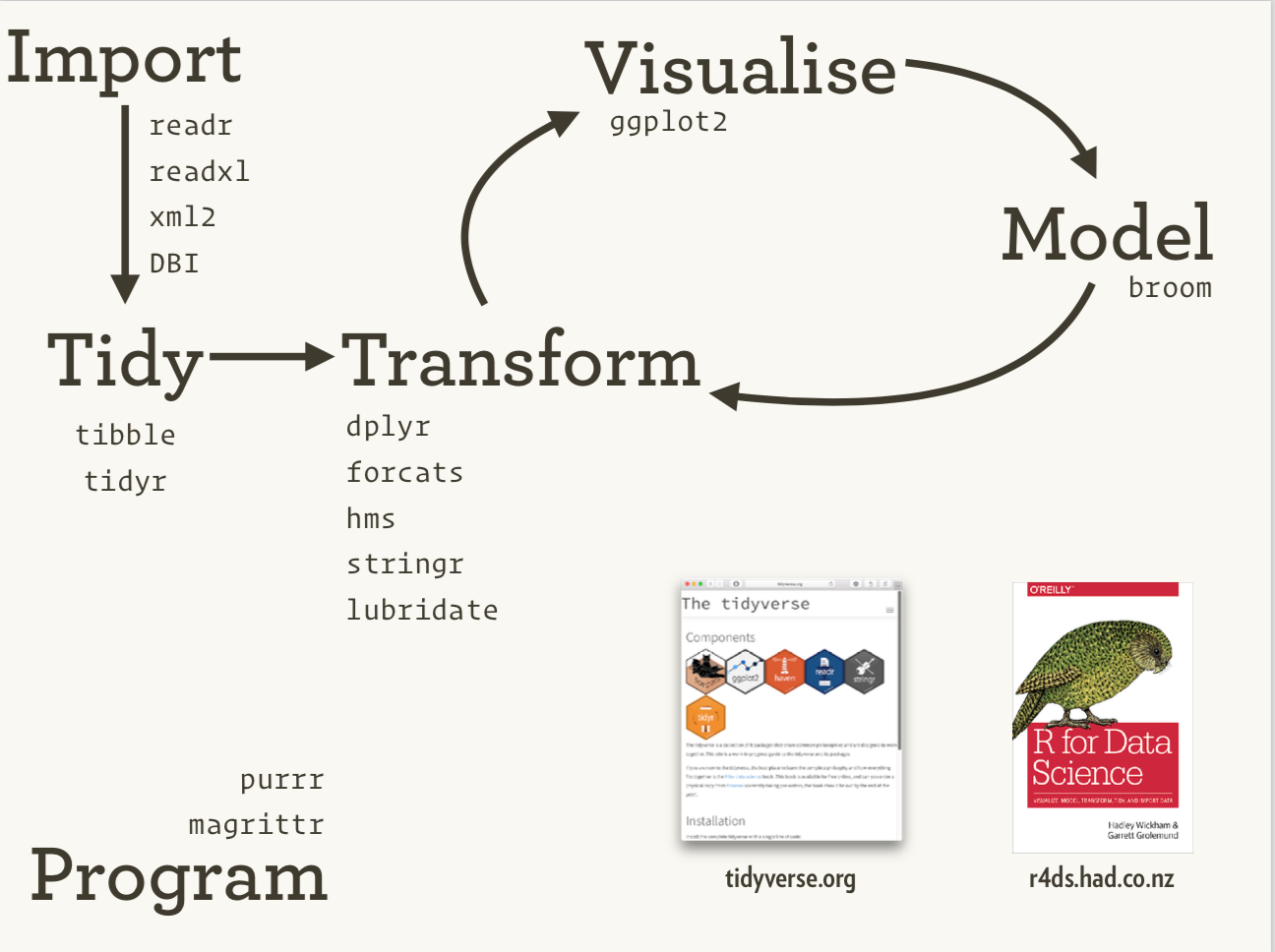
The tidyverse is an ecosystem of R packages for data science, originally conceived by Hadley Wickham.
Our Dataset
We use NHANES data from the training set:
The Pipe Operator
The pipe %>% passes the output of one function as the first argument of the next:
x %>% f(y) is equivalent to f(x, y)Base R also has a native pipe: |>
Why Pipe?
Pipes make code more readable by expressing operations left-to-right:
dplyr: Six Core Verbs
| Verb | Purpose |
|---|---|
filter() |
Select rows by conditions |
select() |
Choose columns |
mutate() |
Create or modify columns |
arrange() |
Sort rows |
summarize() |
Collapse to summary statistics |
group_by() |
Partition data into groups |
filter(): Select Rows
filter(): Multiple Conditions
| PID | BMXBMI | LBXGLU | RIDAGEYR | male |
|---|---|---|---|---|
| 29 | 36.94 | 142.1 | 62 | 1 |
| 211 | 31.30 | 133.7 | 76 | 1 |
| 266 | 29.51 | 165.6 | 82 | 1 |
| 272 | 22.36 | 221.2 | 56 | 1 |
| 695 | 26.10 | 166.1 | 59 | 1 |
select(): Choose Columns
Other helpers: ends_with(), contains(), matches(), one_of()
select(): Example Output
Combining filter and select
mutate(): Create New Columns
case_when(): Recode Variables
case_when works like a switch statement inside mutate:
case_when(): Age Groups
arrange(): Sort Rows
summarize(): Collapse to Summary
group_by(): Partition Data
group_by(): Multiple Groups
| gender | obese | n | mean_glucose |
|---|---|---|---|
| Female | FALSE | 6909 | 94.93 |
| Female | TRUE | 2085 | 105.23 |
| Female | NA | 1796 | 102.56 |
| Male | FALSE | 7028 | 100.74 |
| Male | TRUE | 1450 | 110.94 |
| Male | NA | 1736 | 117.89 |
Tidy Data
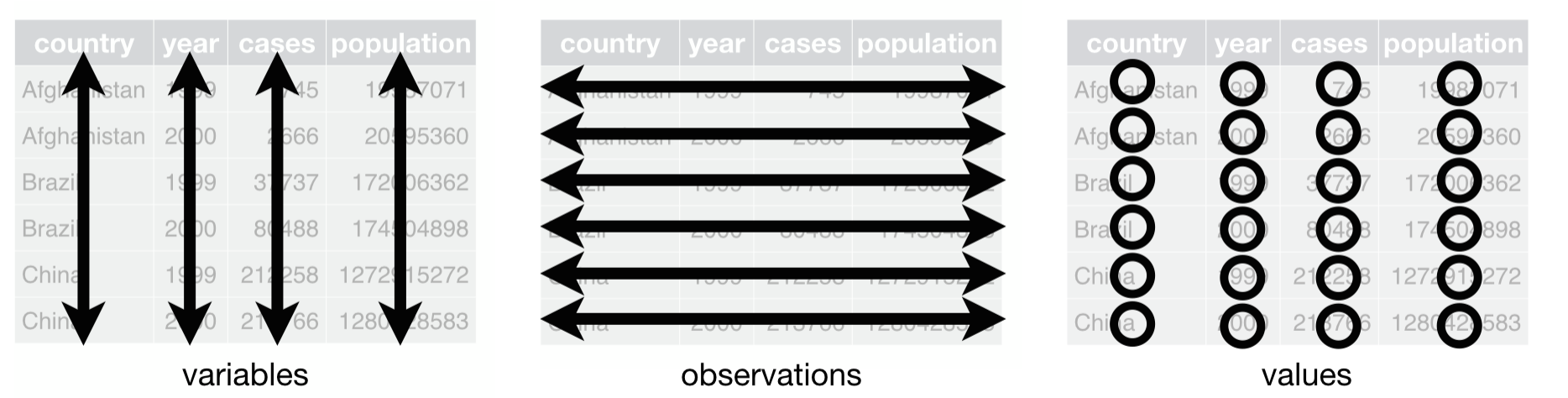
Tidy data principles (Hadley Wickham, inspired by E.F. Codd):
- Each variable forms a column
- Each observation forms a row
- Each type of observational unit forms a table
Is NHANES Tidy?
| PID | BMXBMI | LBXGLU | gender | ethnicity |
|---|---|---|---|---|
| 17050 | 23.57 | NA | Male | Mexican American |
| 1954 | 25.32 | 94.3 | Male | Black |
| 8258 | 15.77 | NA | Male | Other Hispanic |
| 11012 | 20.98 | NA | Female | White |
| 4322 | 37.97 | 97.8 | Female | White |
Yes — each row is one participant’s measurements.
pivot_longer(): Wide to Long
| PID | measurement | value | gender |
|---|---|---|---|
| 1 | BMXBMI | 14.90 | Female |
| 1 | LBXGLU | NA | Female |
| 2 | BMXBMI | 24.90 | Male |
| 2 | LBXGLU | 83.70 | Male |
| 3 | BMXBMI | 17.63 | Female |
| 3 | LBXGLU | NA | Female |
pivot_wider(): Long to Wide
ggplot2: Grammar of Graphics
Components of a ggplot:
data: the data frameaes(): aesthetic mappings (x, y, color, size, shape)geom_*(): geometric objects (points, bars, lines)facet_*(): faceting for small multiplestheme_*(): appearance
Histogram
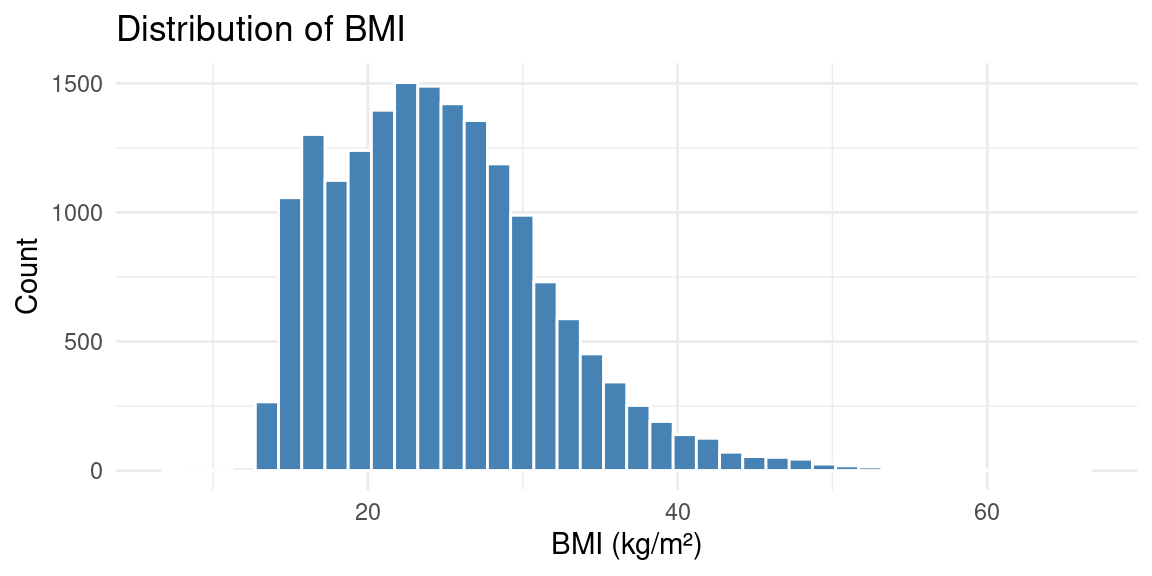
Histogram with Facets
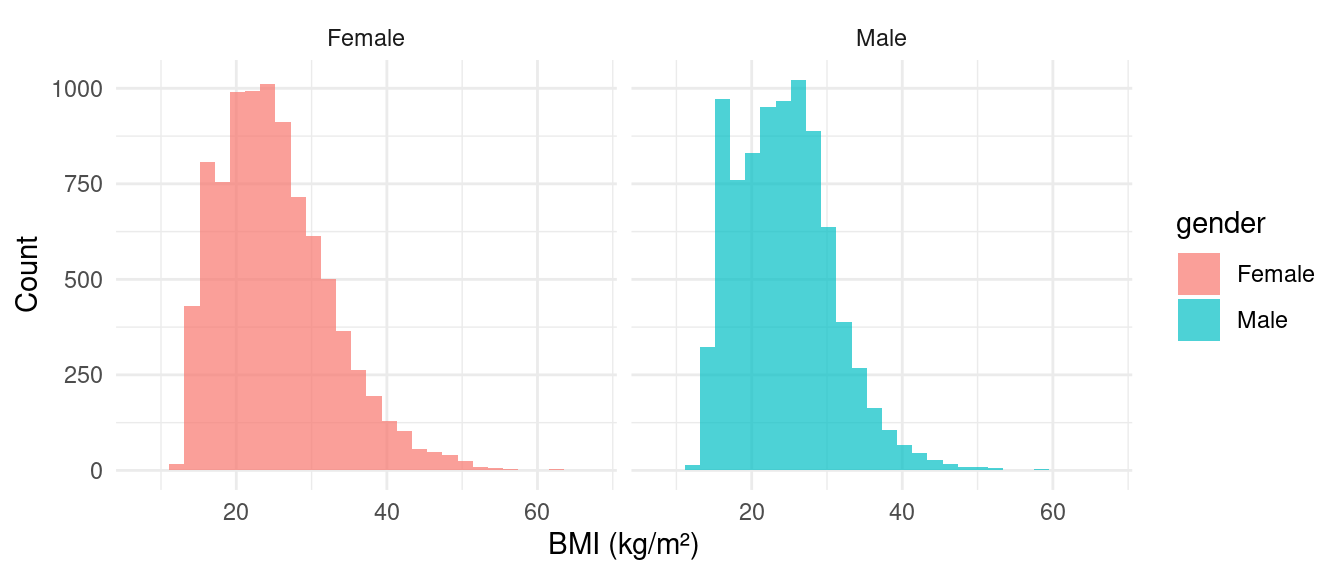
Boxplot
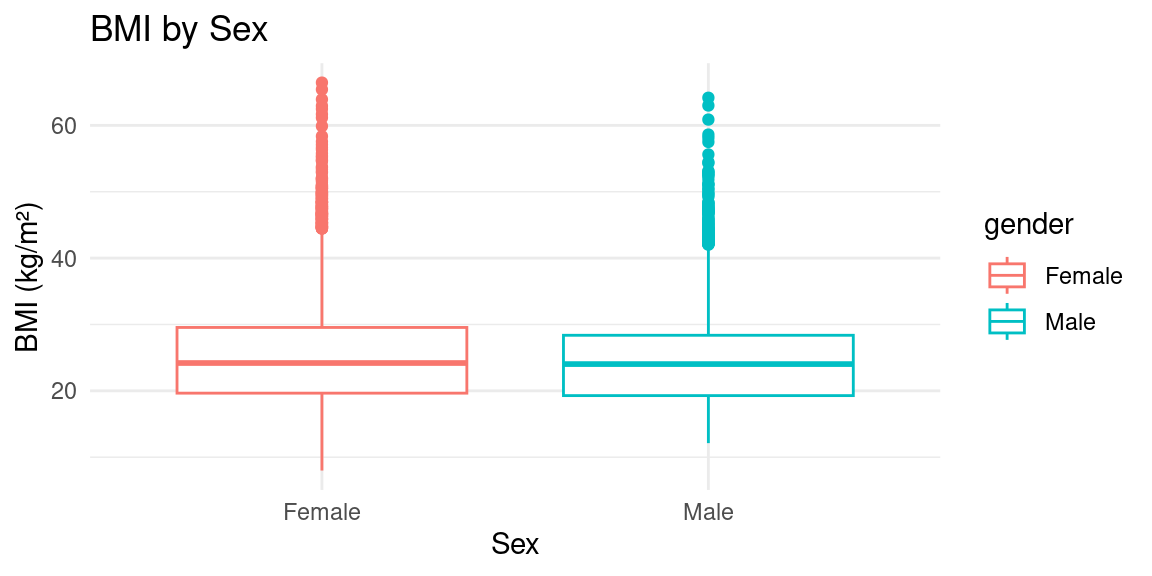
Boxplot with Facets by Ethnicity
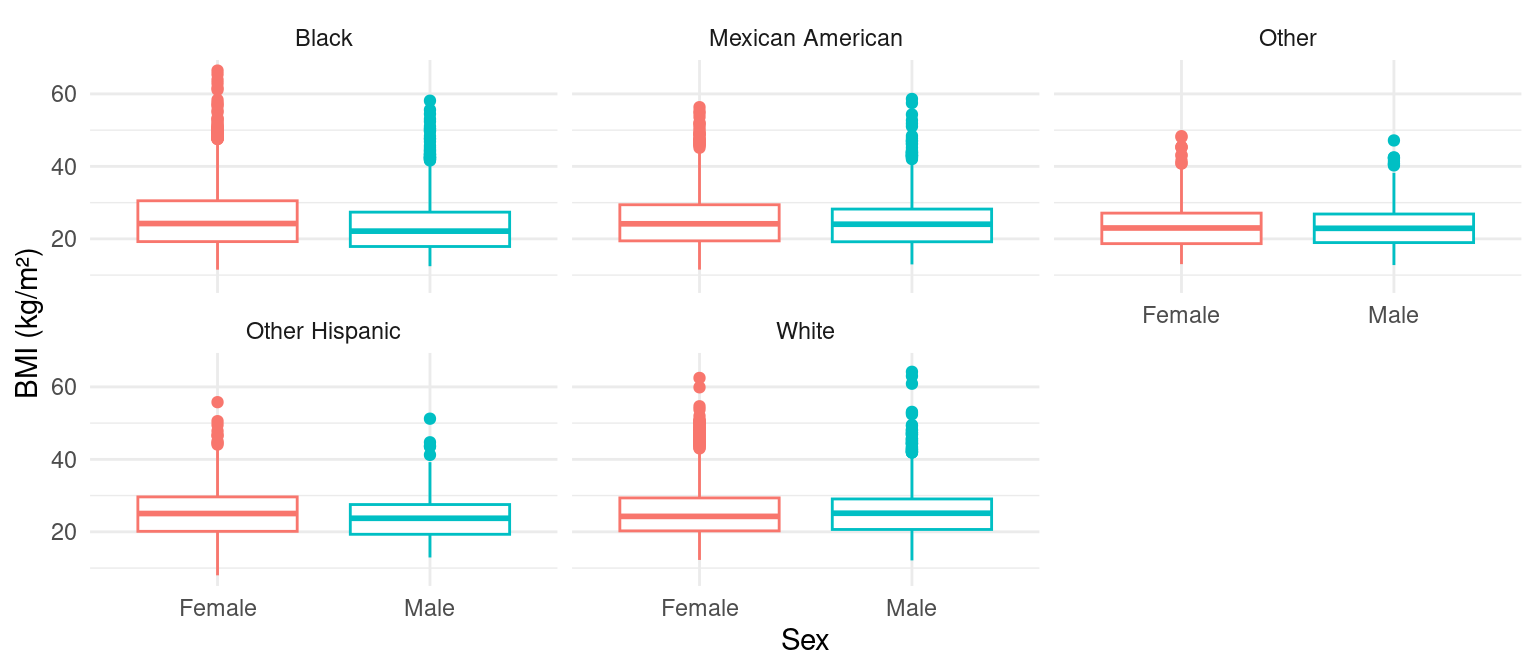
Scatterplot
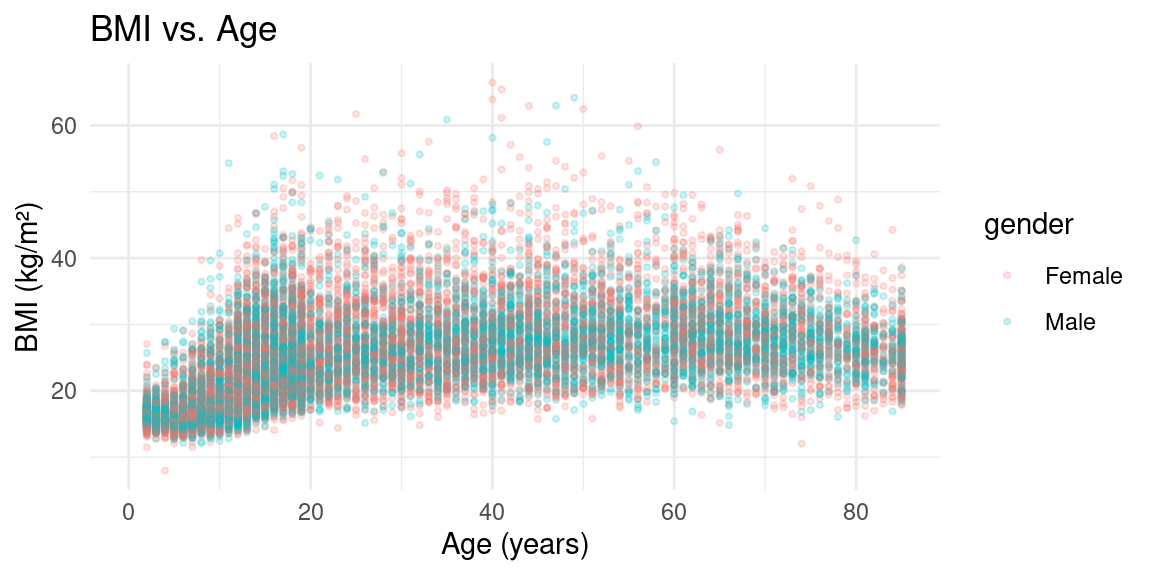
Scatterplot with Facets
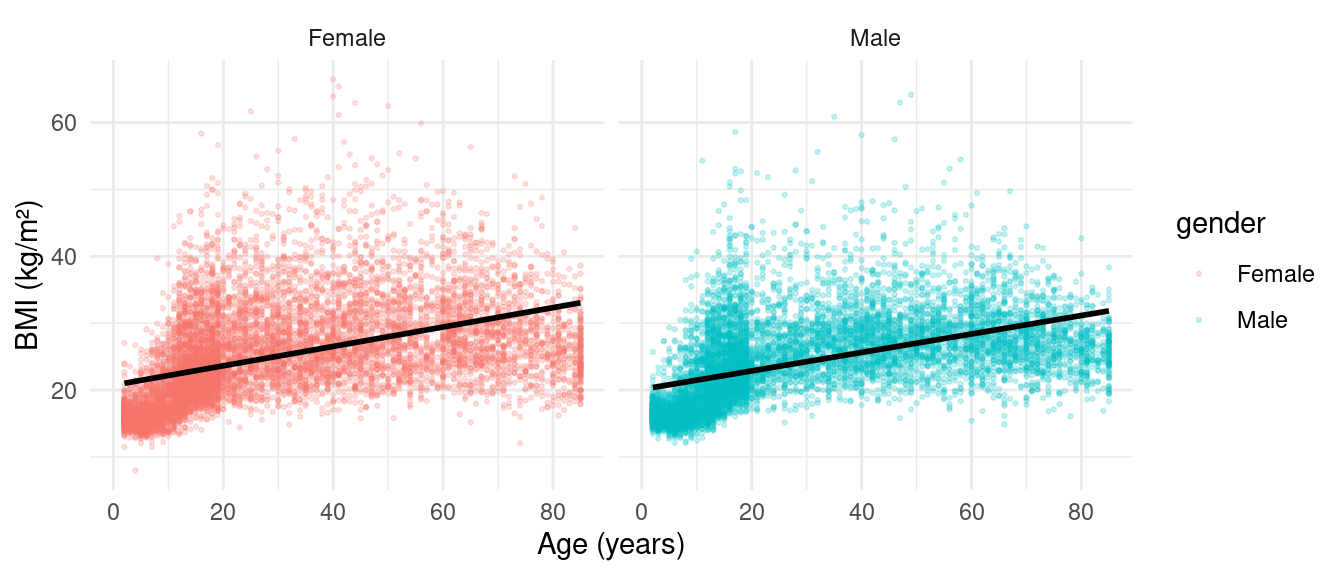
Linear Regression in R
Model the relationship: \(y = \alpha + \sum_{i=1}^{M} \beta_i x_i\)
Three levels of output:
- Model level: \(R^2\), residual standard error
- Term level: coefficient estimates, p-values
- Observation level: predictions, residuals
Model Summary
Call:
lm(formula = LBXGLU ~ BMXBMI + RIDAGEYR + male + black + mexican +
other_hispanic + other_eth, data = nhData)
Residuals:
Min 1Q Median 3Q Max
-77.67 -11.76 -4.21 3.49 592.88
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 64.46384 1.80388 35.736 < 2e-16 ***
BMXBMI 0.57673 0.06243 9.237 < 2e-16 ***
RIDAGEYR 0.39515 0.01863 21.212 < 2e-16 ***
male 5.62365 0.76229 7.377 1.82e-13 ***
black 1.98181 1.02949 1.925 0.05427 .
mexican 5.83894 0.94913 6.152 8.12e-10 ***
other_hispanic 5.56316 1.84842 3.010 0.00263 **
other_eth 5.29182 2.15850 2.452 0.01425 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 30.25 on 6324 degrees of freedom
(14672 observations deleted due to missingness)
Multiple R-squared: 0.1068, Adjusted R-squared: 0.1058
F-statistic: 108 on 7 and 6324 DF, p-value: < 2.2e-16Interpreting the Coefficients
BMXBMI: change in glucose per 1 kg/m² increase in BMIRIDAGEYR: change in glucose per 1 year increase in agemale: difference in glucose for males vs. females (reference)black,mexican, etc.: difference vs. white (reference group)
Reference categories: female for sex, white for race/ethnicity.
The broom Package
broom tidies model output into data frames:
| Function | Returns | Description |
|---|---|---|
glance() |
1-row tibble | Model-level summary (\(R^2\), AIC) |
tidy() |
Per-term tibble | Coefficients, p-values per variable |
augment() |
Per-observation tibble | Predictions, residuals |
glance(): Model-Level Summary
tidy(): Term-Level Summary
| term | estimate | std.error | p.value |
|---|---|---|---|
| (Intercept) | 64.4638 | 1.8039 | 0.0000 |
| BMXBMI | 0.5767 | 0.0624 | 0.0000 |
| RIDAGEYR | 0.3952 | 0.0186 | 0.0000 |
| male | 5.6237 | 0.7623 | 0.0000 |
| black | 1.9818 | 1.0295 | 0.0543 |
| mexican | 5.8389 | 0.9491 | 0.0000 |
| other_hispanic | 5.5632 | 1.8484 | 0.0026 |
| other_eth | 5.2918 | 2.1585 | 0.0142 |
augment(): Observation-Level
Running Many Regressions with nest_by
Goal: Predict BMI and glucose as a function of age, sex, ethnicity — stratified by survey year.
Step 1: Make data long
Step 2: nest_by and Fit Models
Step 3: Extract R² from Each Model
Step 4: Extract Coefficients from Each Model
| outcome | SDDSRVYR | term | estimate | p.value |
|---|---|---|---|---|
| BMXBMI | 1 | (Intercept) | 20.075 | 0.000 |
| BMXBMI | 1 | RIDAGEYR | 0.145 | 0.000 |
| BMXBMI | 1 | male | -1.063 | 0.000 |
| BMXBMI | 1 | black | 1.453 | 0.000 |
| BMXBMI | 1 | mexican | 1.209 | 0.000 |
| BMXBMI | 1 | other_hispanic | 0.854 | 0.005 |
| BMXBMI | 1 | other_eth | 0.818 | 0.028 |
| BMXBMI | 2 | (Intercept) | 19.948 | 0.000 |
| BMXBMI | 2 | RIDAGEYR | 0.149 | 0.000 |
| BMXBMI | 2 | male | -0.632 | 0.000 |
Putting It All Together
The tidyverse workflow for ExWAS:
- Load and clean data with dplyr (
mutate,filter,select) - Reshape data with tidyr (
pivot_longer) - Visualize data with ggplot2
- Model with
lm()orsvyglm() - Tidy results with broom (
tidy,glance) - Scale analyses with
nest_by+map
Summary
- The pipe operator makes code readable and composable
- dplyr provides 6 core verbs for data manipulation
- case_when enables flexible variable recoding
- pivot_longer/wider reshapes data between wide and long formats
- ggplot2 creates publication-quality visualizations
- broom tidies model output for downstream analysis
- nest_by scales regressions across groups
What’s Next?
Module 3: Statistical Foundations for ExWAS
- Survey sampling and svydesign/svyglm
- Log transformation and standardization
- Multiple testing correction
- Confounding and DAGs
Supported By
This course is supported by the National Institutes of Health (NIH):
- National Institute of Environmental Health Sciences (NIEHS): R01ES032470, U24ES036819
- National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK): R01DK137993